Vision Statement
The
Virtual Genetics Lab (VGLII) is a simulation of transmission genetics that
approximates, as closely as possible, the hypothesis-testing environment of
genetics research. In this lab, students cross hypothetical creatures
and examine the progeny in order to determine the mechanism of inheritance of
a particular trait. As in actual research, it is not possible to 'see
the answer' - the student must decide for herself when she has collected
enough data to be sure of her model. The goal is to have students
understand the logic of genetic analysis - how one can use crosses to
determine how a trait is inherited and reinforce their understanding of
transmission genetics.
VGLII also allows students to enter their answers - their model of how
their traits are inherited - in a form that allows their work to be
automatically graded by their instructor. Click the GRADING button
to the left for details.
Issues with Java? (2024-10-26)
I've heard from some users that Java is no longer available on their computers. Since Java is essential for VGL, I have created a non-java version of VGLII that runs in a web browser. I have posted a version of this program, jsVGL, for users to test and give me feedback. jsVGL can load and save work, so it can be used in many of the most common applications of VGLII. If you're interested, please give it a try and let me know what you think.In addition, some versions of Mas OSX give a popup that says "Unable to Load Java Runtime Environment" when you try to run VGL. The work-around is to:
- Right-click the VGLII icon and choose "Show Package Contents"
- In the Finder window that appears, open "Contents"
- Open the "Java" folder
- In that, double-click the "VGLII.jar" file.
What's new in Version 3?
In response to requests from the user community, I have added several features to VGLII including:- Chi-Squared Tests: The Summary Chart now includes the ability to calculate χ2 values.
- Epistasis and Complementation: models can optionally include these gene interactions.
- Super Cross: students can carry out a cross with up to 2000 offspring for getting statistically-significant ratios.
- Model Builder: students can enter and save their developing genetic models.
- Grading Option: instructors can compare students' answers in the Model Builder with the correct answers and grade students work. Students can also add "comments to the instructor" along with their answers; these comments can then be viewed by the grader.
VGLII Videos
I have made two short videos demonstrating VGLII: VGLII in about 1 minute. This shows how VGLII works. (Click the image to play the movie)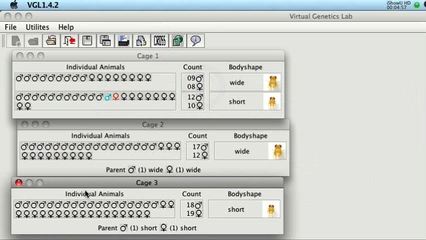
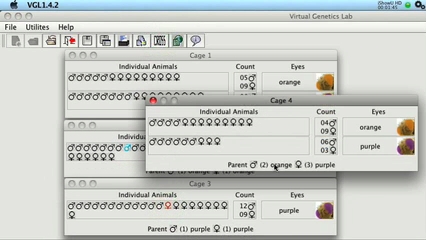
VGLII in more detail. This is about 5 minutes long
and gives more detail on VGLII and how to use it. (Click the image to play the movie)
VGLII Handout
I made this handout describing VGLII and including a sample student lab report.Click the image to see it in full size.

A Sample VGLII Lab Report
One of Professor Sophie Javerzat's students prepared this outstanding report based on a very complex VGLII problem.VGLII Philosophy and Mission
This program is based on the Genetics Construction Kit (GCK) developed by
BioQUEST. The original GCK was written in the
mid-1980's and ran on the Macintosh Classic operating environment. VGLII is designed
as a newer version of GCK that is compatable with Windows and Unix-based operating
systems and that takes advantage of the improved user interface available on
newer computers. VGLII includes all features of GCK and then some.
A new version of GCK is expected soon; check the
BioQUEST website for updates. A comparison
of some key features of VGLII and GCK is
shown at the bottom of this page. VGLII is a revised version of VGL, which is no longer
being maintained. VGLII can do all that VGL could and more!
In order that the exercise is based on the logic of genetic analysis rather than student's ability to search the internet, the creature under study is hypothetical, the traits are assigned randomly, and the mode of inheritance differs with each run of the program. Traits can be either autosomal or sex-linked (with XX females and XY males or with ZZ males and ZW females) and simply dominant or co/incompletely dominant. The form of the trait which is dominant, recessive or seen in the heterozygote will also be determined randomly. The instructor can customize the program to limit the range of possibilities that the students could face. In this way, students can be gradually introduced to problems of increasing complexity.
In a real genetics lab, students have to deal with real-world creatures that die, fail to mate, or escape. In addition, most creatures have generation times of several weeks or months, thus limiting the number of crosses an individual can perform during a semester. Although these real-world constraints have educational value, the point of the VGLII is to understand the logic of genetic analysis rather than the technical details. The VGLII allows the students to maximize the number of crosses they can perform, thus maximizing their chance to understand the underlying analysis.
In this lab, students have to decide:
-
What are the possible modes of inheritance.
-
Which organisms to cross.
-
How to interpret their data.
-
When they have enough data to be sure of their conclusions.
In order to perform a cross, the students select male and female parents of their choice from any generation and click the "Cross" button. The program keeps track of the genotypes of the parents and generates their offspring probabilistically. In that way, the phenotypic ratios of the offspring will approximate, but not necessarily be identical to the classic 1:1, 1:2:1, and 1:3 ratios. The students have to decide for themselves whether their data are consistent or inconsistent with their expected ratios. While some crosses will generate useful information, others will be uninformative; the students will have to choose what to cross and what to make of the results.
Cross results are presented in a table sorted by phenotype with a display of the numbers of male and female individuals of each phenotype. In addition, a thumbnail image in the table gives a visual display of the phenotype.
Because of this freedom to choose what to cross, different student groups will adopt different strategies for analysis. Some will plan each cross carefully and analyze the results as they go along while others will do many haphazard crosses and try to sort out the results later. VGLII thus accommodates the wide range of successful strategies and does not limit the students to one particular pattern.
Although VGLII is designed so that students cannot 'look behind the scenes' to see the underlying model or the genotypes of individuals, the instructor can generate problems in 'practice mode'. In this mode, students are able to click on individuals and see their genotypes as well as the underlying genetic model. This mode is designed for the beginning students to help familiarize them with the program and the most basic genetic analysis.
Finally, the program has several important software features:
-
it is extremely robust and reliable since it will be used by beginning students
-
it is compatible with both Mac, UNIX/Linux, and PC
-
it is disseminated by the www
-
it has an easy-to-use color GUI
VGLII has been used by over 3500 biology students at UMass Boston since 2003.
VGLII is in use in many countries including the US, France, and Korea.
VGLII was designed and developed by a team of four Master's degree students in Computer Science at UMass Boston and Brian White.
A comparison of VGLII and GCK is shown below.
Comparison of VGLII and GCK
| Program | Compatible OSs | Genes per problem | Sex-linkage | User Interface | Phenotype Display | Answer Submission | Price |
| VGLII |
-Win98 & up -UNIX/Linux -Mac OS 10.1 through 10.6 |
1 to 3 only | XX/XY & ZW/ZZ | Color | Color Images | Possible | Free |
| GCK | Mac OS 6 thru 10.3 | 1 to 3 | XX/XY only | Black & white | Text only | Not possible | Nominal charge |